A New Lens on Biological Complexity
For decades, biology relied on bulk sequencing, averaging signals across millions of cells. While powerful, this approach masked the extraordinary diversity that exists within tissues. Single‑cell genomics changed that paradigm entirely. By sequencing individual cells, researchers can now uncover hidden subpopulations, rare cell states, and dynamic transitions that shape development, immunity, and disease (Cuomo et al., 2023). This shift from averages to single‑cell resolution has become one of the most influential revolutions in modern genomics.
Revealing the Hidden Architecture of Tissues
Single‑cell sequencing has shown that no tissue is truly uniform. The immune system, once thought to consist of a handful of cell types, is now understood as a vast landscape of finely tuned subpopulations. Tumors, previously classified by broad histological categories, reveal intricate mosaics of malignant clones, stromal cells, and immune infiltrates. Even the brain — the most complex organ — is being mapped cell by cell, uncovering previously unknown neuronal subtypes and developmental trajectories (Wu et al., 2024). These insights are reshaping how we diagnose disease and design targeted therapies.
From Microbial Dark Matter to Human Health
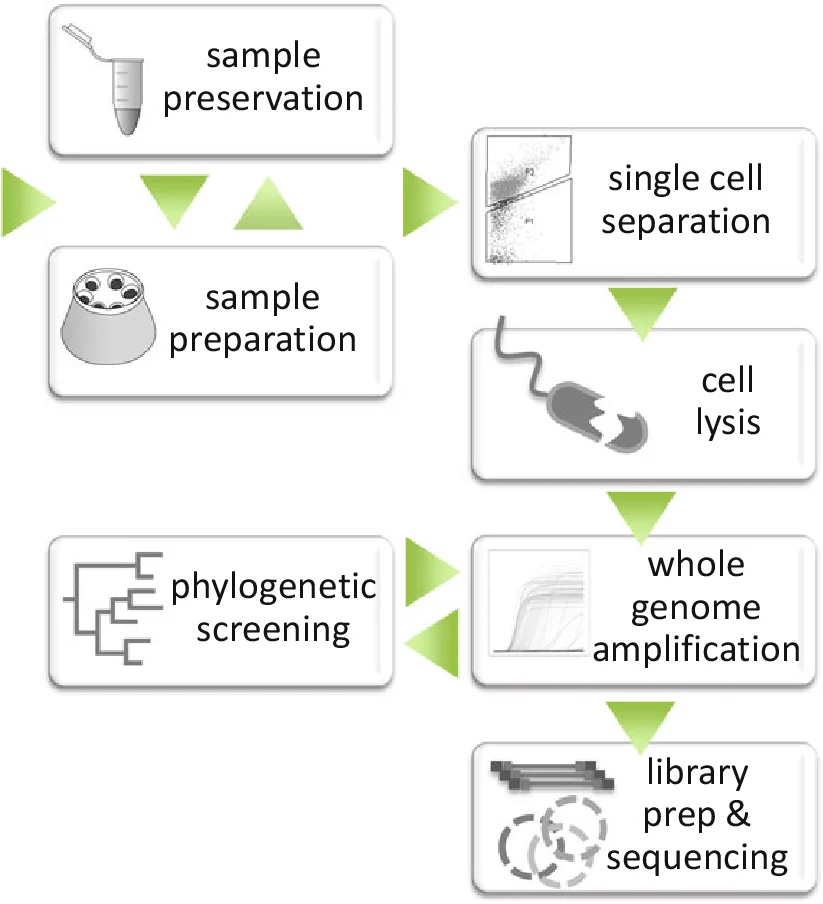
Early single‑cell genomics was applied to environmental microbiology, enabling the sequencing of uncultured microorganisms — the so‑called “microbial dark matter.” Today, the technology has expanded into nearly every biomedical field. In oncology, single‑cell RNA‑seq identifies resistant clones before treatment begins. In immunology, it maps T‑cell exhaustion and vaccine responses with unprecedented precision. In developmental biology, it reconstructs lineage trees that reveal how a single fertilized egg becomes a complex organism (Cuomo et al., 2023). The breadth of applications continues to grow as methods become more scalable and affordable.
The Rise of Spatial and Multimodal Single‑Cell Technologies
The latest wave of innovation goes beyond sequencing alone. Spatial transcriptomics now preserves the physical location of each cell within a tissue, allowing researchers to see not only what a cell expresses but where it lives and whom it interacts with. Multimodal platforms combine RNA, chromatin accessibility, protein markers, and even CRISPR perturbations within the same cell (De Jonghe et al., 2024). These integrated approaches are transforming single‑cell genomics from a descriptive tool into a mechanistic engine for discovery.
A Foundation for Precision Medicine
As datasets grow to millions of cells, single‑cell genomics is becoming a cornerstone of precision medicine. It enables clinicians to identify patient‑specific disease signatures, predict therapeutic responses, and monitor treatment at cellular resolution. The future points toward real‑time single‑cell diagnostics, where clinicians can track disease evolution as it happens. What began as a niche research technique is rapidly becoming an essential component of next‑generation healthcare (Wu et al., 2024).
References
Cuomo, A. S. E. et al. (2023). Single‑cell genomics meets human genetics. Nature Reviews Genetics.
Wu, X. et al. (2024). Single‑cell sequencing to multi‑omics: technologies and applications. Biomarker Research.
De Jonghe, J. et al. (2024). scTrends: A living review of commercial single‑cell and spatial ’omic technologies. Cell Genomics.
Image source: https://www.researchgate.net/publication/328140505_Single-Cell_Genomics_of_Microbial_Dark_Matter_Methods_and_Protocols